The study investigates the impact of foreign remittance and economic complexity on carbon () emissions by using the data for Asian countries over the period of 1
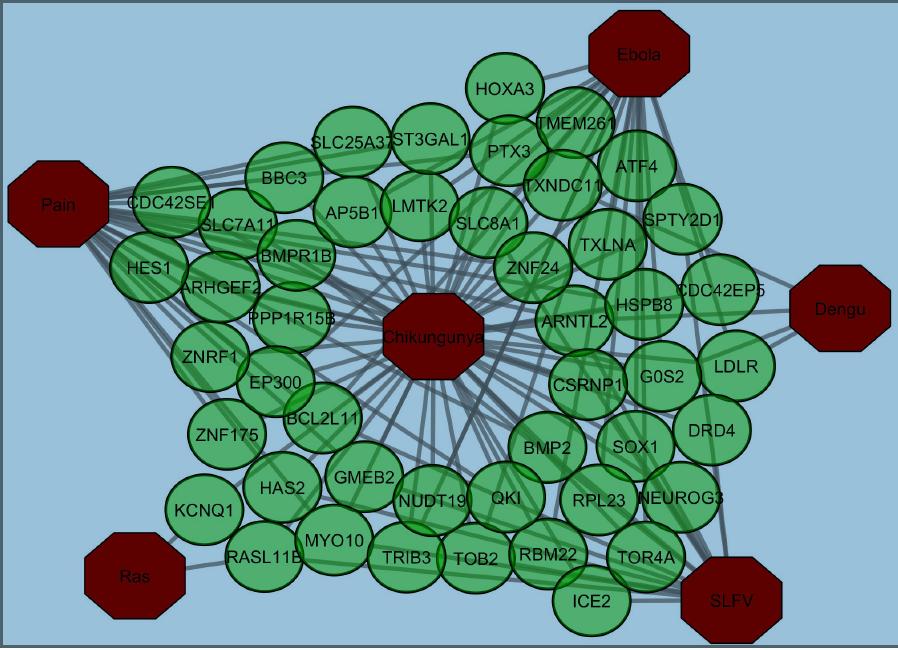
The Chikungunya (CKV) viral infection is a global health burden characterized by the neurologic complications with CKV infection. The present study aimed to discover molecular signatures for comorbidity of viral infections. We have analyzed transcriptome (RNA-Seq data and microarray) from Chikungunya, Ebola, Dengu, Semlikhi Forest Virus (SLFV), Pain, Rash, and control datasets. Identifying the common genes among the above diseases, we constructed relationship networks centered on CHIKV virus. After that, we also constructed protein protein interaction network (PPI) considering the DEGs of CHKIV and identified hub genes based on topological analysis. A total 500 differentially expressed genes (DEGs) was identified associated with CHIKV infections. It was also found that 105 genes were common in both CHIKV and ebola infections. However, CHIKV shared under 24 significant transcripts with other alphaviruse infections. We also found that 49 genes shared with pain and 10 genes with rash. We identified from infectome-diseasome analysis that chikungunya virus infection is mostly related with the ebola virus infection. But from phylogenic analysis, it was found that these virus infections are related. The pathway and ontology analysis revealed CHIKV infection most resembles ebola fever.